Advantages:
- Therapeutic Targets: Identified essential genes in Pa aggregate formation that can be potentially targeted with therapeutics
- Improved Infection Model: use of synthetic CF sputum media that closely mimics the physical and nutritional environment of CF sputum
- Potential for Broader Applications: can be extended to other bacterial species and infection models
Summary:
Chronic lung infections cystic fibrosis (CF) caused by Pseudomonas aeruginosa (Pa) are difficult to treat due to the formation of small bacterial aggregates. Unlike traditional biofilms, the smaller aggregates present unique challenges and are more persistent and resistant to treatment. Their formation mechanisms and genetic underpinnings remain poorly understood. Identifying the genes and pathways specific to these aggregates is crucial for designing targeted therapies that disrupt their formation and mitigate chronic infection.
Our researchers identified genes essential for the formation of Pseudomonas aeruginosa (Pa) aggregates in cystic fibrosis (CF) sputum. By combining RNA sequencing with high-resolution microscopy and predictive modeling, they identified 13 genes that are differentially expressed during the formation of Pa aggregates in synthetic CF sputum media (SCFM2). Predictive modeling linked the genes to metabolic shifts, iron acquisition, surface modification, and quorum sensing. Analysis of spatial phenotypes of the PA mutants carrying the disrupted genes revealed a subset of the genes, where mutations resulted in loosely packed aggregates and slower growth rates, indicating that these genes are crucial for maintaining aggregate integrity and density.
This study provides a foundation for the understanding of the genetic and biological mechanisms underlying bacterial aggregation and offers novel potential therapeutic targets for Pseudomonas aeruginosa treatment.
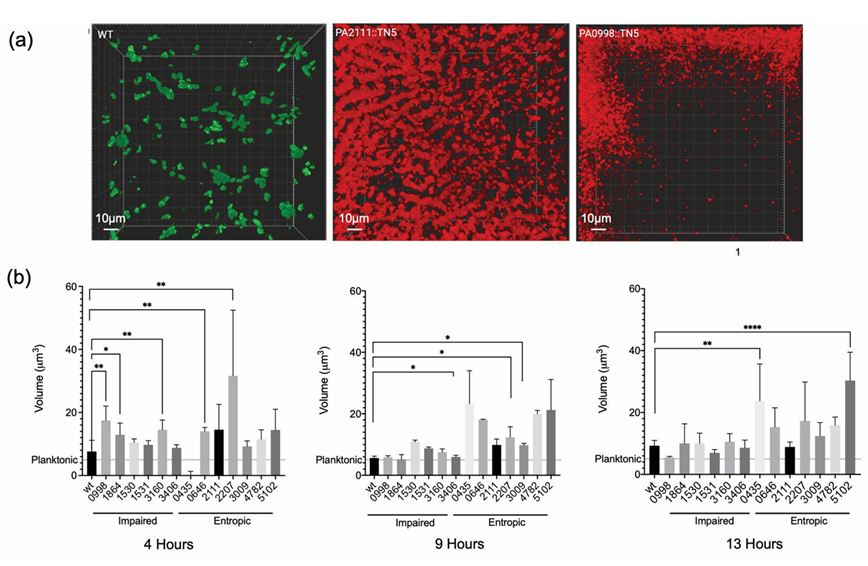
Figure: Pseudomonas aeruginosa Mutant Aggregate Phenotypes
(a) Examples of aggregation, with WT shown in green and two representative mutants shown in red. (b) Average aggregate volume of transposon mutants over time.
Desired Partnerships:
- License
- Sponsored Research
- Co-Development